- Title
-
Deletion of Slc1a4 Suppresses Single Mauthner Cell Axon Regeneration In Vivo through Growth-Associated Protein 43
- Authors
- Li, K., Fan, D., Zhou, J., Zhao, Z., Han, A., Song, Z., Tang, X., Hu, B.
- Source
- Full text @ Int. J. Mol. Sci.
|
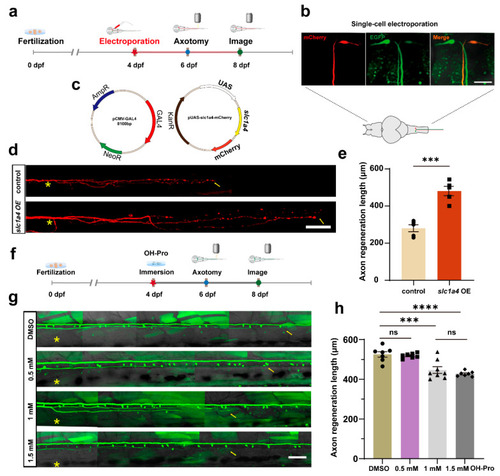
Overexpression of slc1a4 in Mauthner cells promotes axon regeneration in vivo. ( |
|
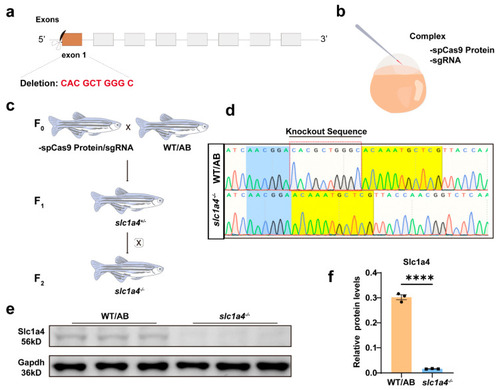
Identification of |
|
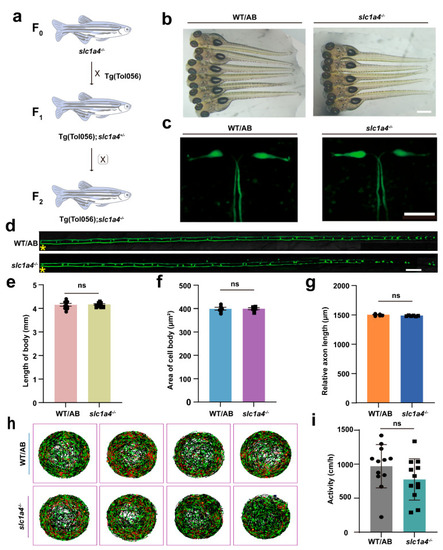
Deficiency in Slc1a4 does not affect development of M-cells and motor function of juvenile fish. ( |
|
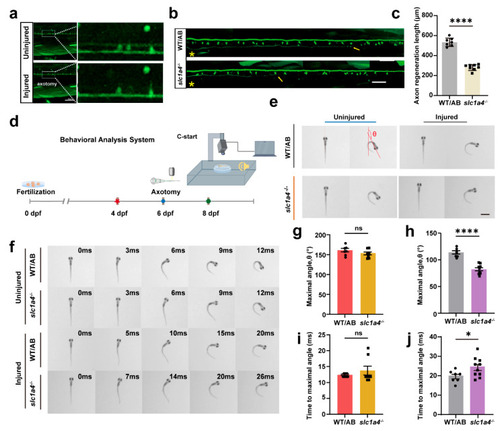
Deficiency in Slc1a4 suppresses Mauthner cell axon regeneration and relative function in vivo. ( |
|
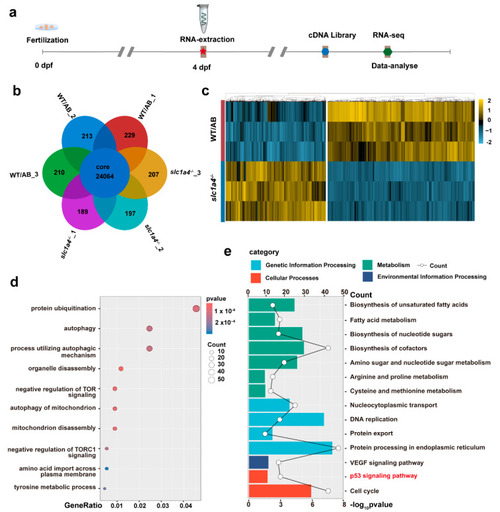
RNA-seq revealed that Slc1a4 may influence regeneration of Mauthner axons through P53 signal pathway. ( |
|
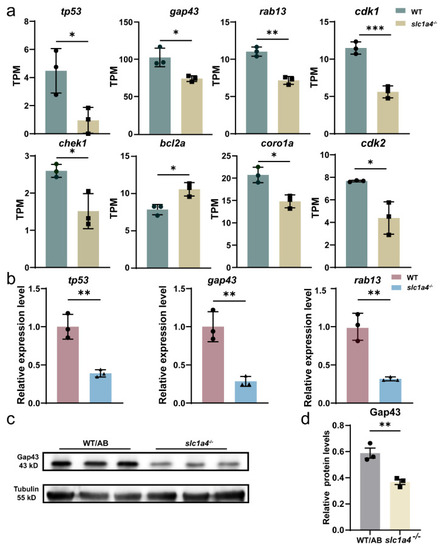
Slc1a4 deletion may inhibit Mauthner axons’ regeneration via Gap43 suppression. ( |