- Title
-
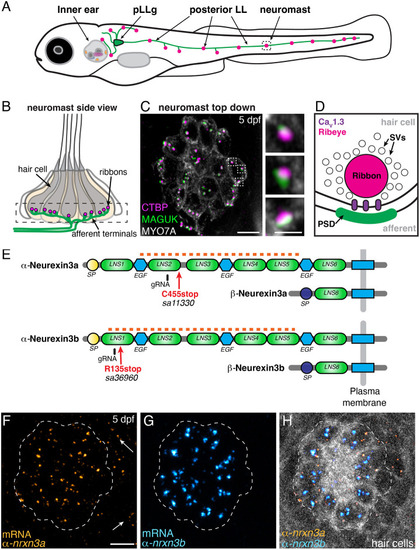
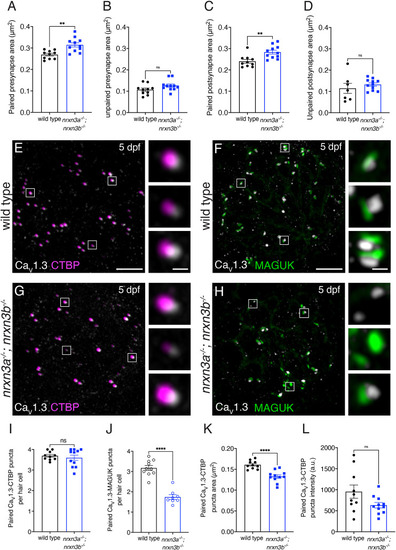
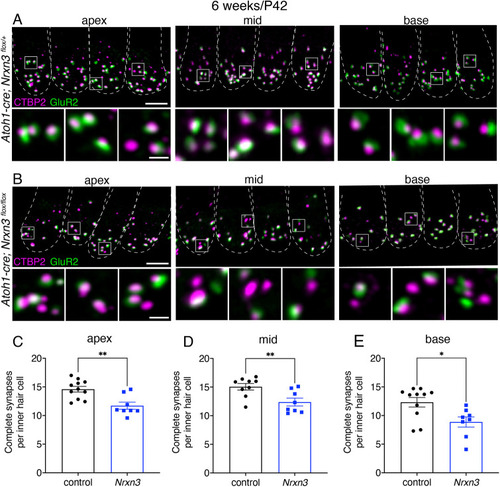
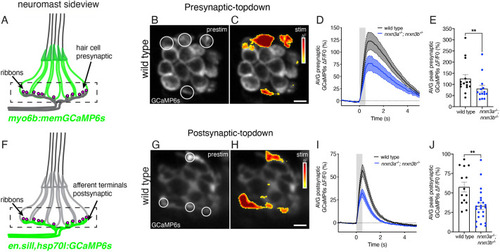
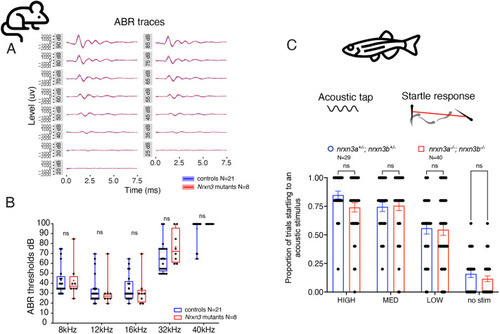
Presynaptic Nrxn3 is essential for ribbon-synapse maturation in hair cells
- Authors
- Jukic, A., Lei, Z., Cebul, E.R., Pinter, K., Tadesse, Y., Jarysta, A., David, S., Mosqueda, N., Tarchini, B., Kindt, K.
- Source
- Full text @ Development
|
EXPRESSION / LABELING:
|
|
|
|
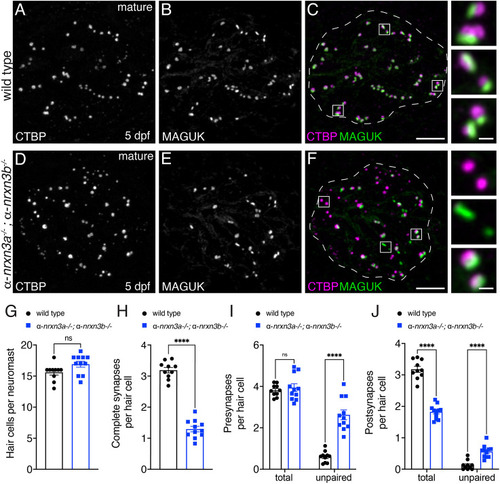
PHENOTYPE:
|
|
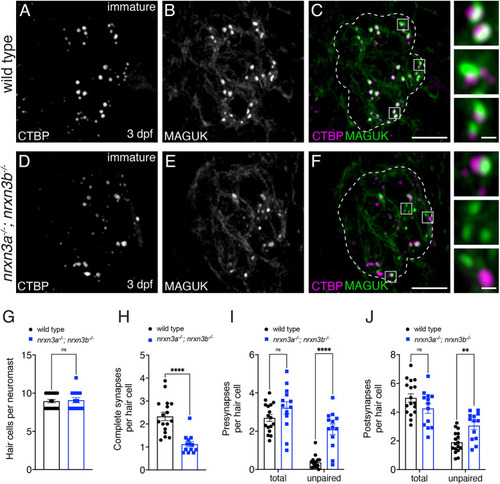
PHENOTYPE:
|
|
|
|
|
|
|
|
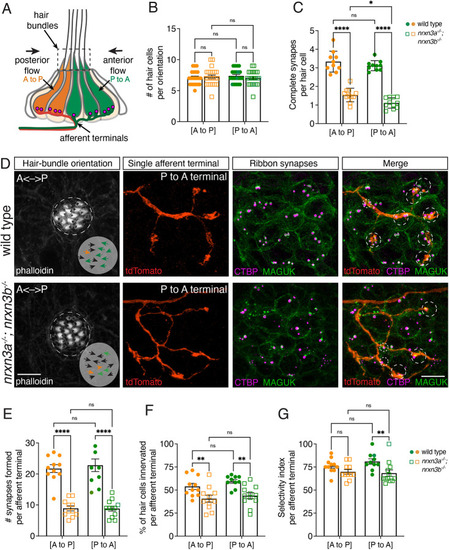
PHENOTYPE:
|