- Title
-
Rare homozygous cilia gene variants identified in consanguineous congenital heart disease patients
- Authors
- Baird, D.A., Mubeen, H., Doganli, C., Miltenburg, J.B., Thomsen, O.K., Ali, Z., Naveed, T., Rehman, A.U., Baig, S.M., Christensen, S.T., Farooq, M., Larsen, L.A.
- Source
- Full text @ Hum. Genet.
|
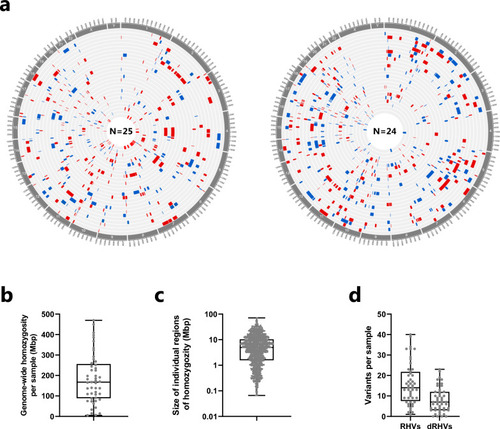
Distribution of rare homozygous variants in CHD patients from consanguineous families. |
|
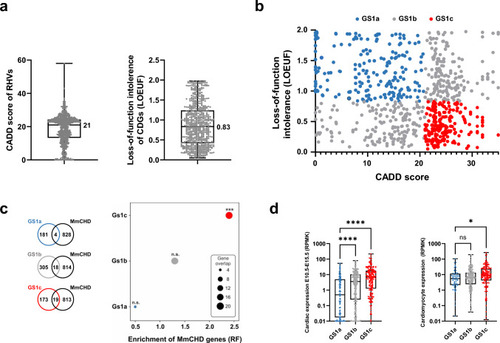
Prioritization of candidate disease genes. |
|
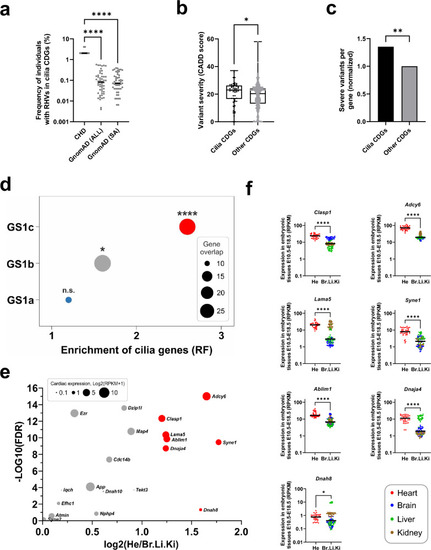
Cilia genes are enriched for rare homozygous variants. |
|
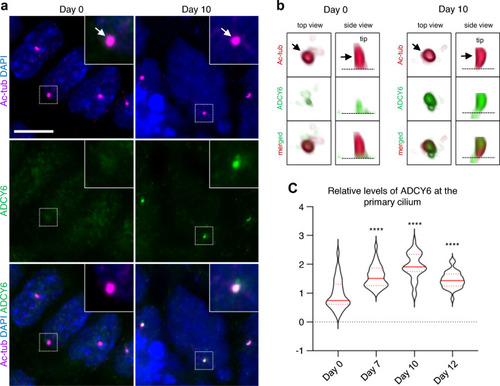
Temporal localization of ADCY6 to primary cilia during cardiomyogenesis. ADCY6 accumulate at primary cilia during differentiation of P19.CL6 cells into cardiomyocytes. |
|
Knock-out of ADCY6 cause heart defects in zebrafish. |