FIGURE 6
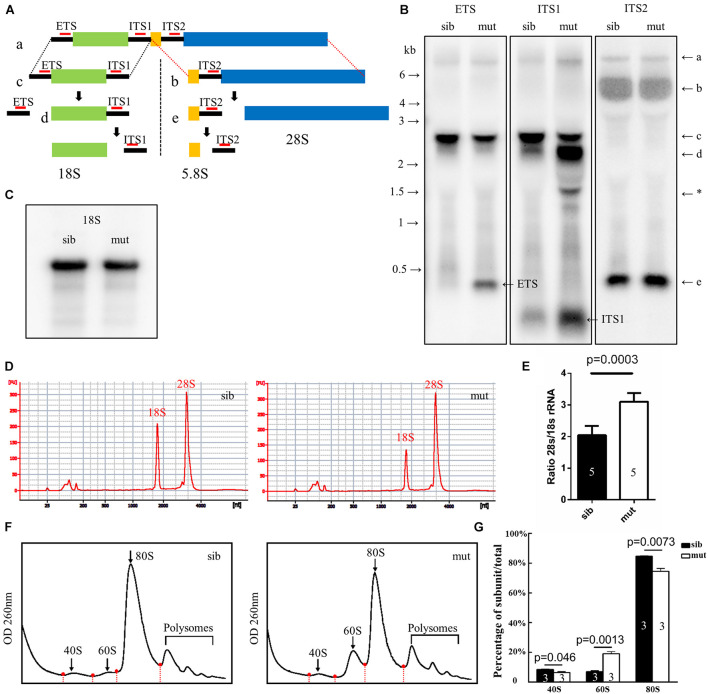
Figure 6. Defects in 18S rRNA processing in ltv1?14/?14 mutant. (A) Schematic diagram showing the processing pathway of 28S, 18S, and 5.8S rRNAs. The hybridization sites of each probe are indicated by red bars, respectively. (B,C) Northern blot analysis using the corresponding probes as indicated to detect 18S rRNA or the intermediate products of rRNA processing. Asterisk: unidentified rRNA intermediate product. (D) Representative analysis result by E-bioanalyzer. (E) The ratio of 28S/18S rRNA is increased in ltv1?14/?14 mutants (N = 5), as compared with siblings (N = 5). Bars represent means with SD. ETS, external transcribed spacer; ITS, internal transcribed spacer. (F) Representative ribosome fractionation results of siblings and ltv1?14/?14 mutants at 4 dpf. The peaks of 40S, 60S, and 80S are indicated by arrows. Red dots on the curves represent the lowest points flanking the peaks. The area under respective peaks of 40S, 60S, and 80S, circled by curves, dashed lines, and baselines are measured. (G) Percentages of 40S, 60S, and 80S in total lysate in siblings and ltv1?14/?14 mutants at 4 dpf. Bars represent means with SD. Quantifications of three independent experiments are analyzed.