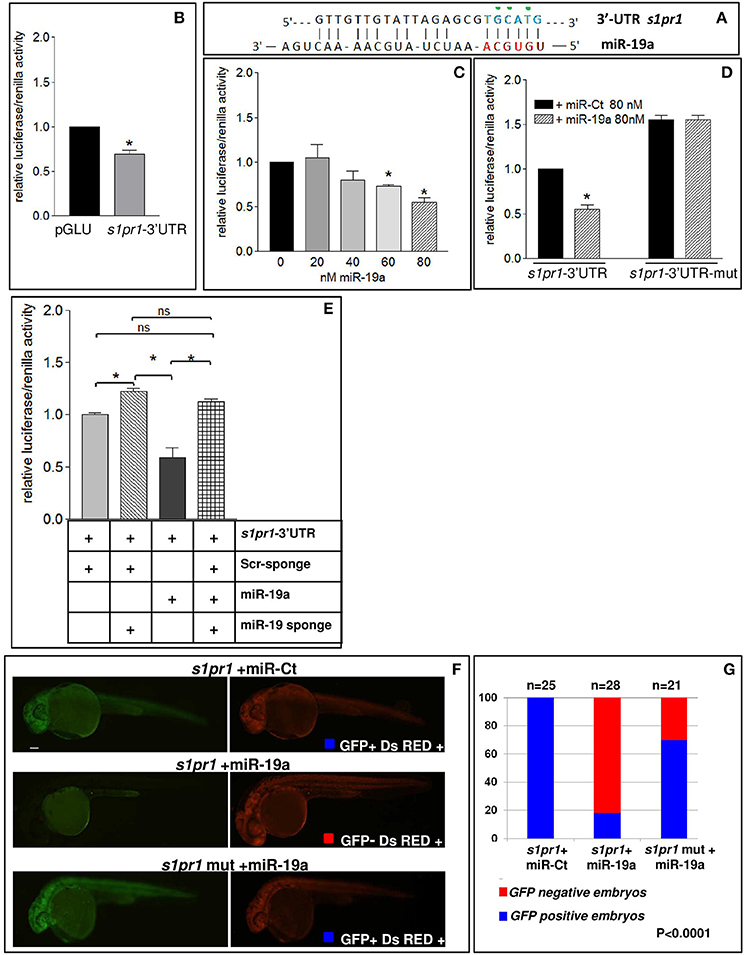
Fig. 4
s1pr1 is a direct target of miR-19a. (A) The computational alignment of a potential target site of miR-19a in s1pr1 of Danio rerio. The green dots indicate the bases mutated to destroy the seed sequence binding in the s1pr1-3?UTRmut construct. (B?E) The 3?UTR of dre-s1pr1 was cloned in the pGLU Dual-luciferase reporter plasmid (s1pr1-3?UTR) and transiently transfected into HK293 cells together with the Renilla luciferase pRL?TK vector as internal standard. The luciferase activity of s1pr1-3?UTR was compared: (B) with the luciferase activity of the empty vector; (C) in the presence of increasing amounts of miR-19a mimic; (D) with the luciferase activity of s1pr1-3?UTRmut (containing the s1pr1 3?UTR mutated in the seed match for miR-19a) in the presence of 80 nM of miR-19a mimic; (E) in the presence of 80 nM of miR-19a mimic and 500 pg of a sponge for miR-19a or 500 pg of a scrambled sponge. In each transfection the total amount of the transfected miRNA was kept constant by adding a scrambled miRNA (miR-Ct) to the specific miRNA to obtain 80 ng. *p < 0.05. (F,G) miR-19a negatively regulates the sensor s1pr1-GFP containing the wt3?UTR of s1pr1 downstream the GFP sequence but it is inefficient on the mut s1pr1-GFP; (F) representative 24 hpf embryos injected with wtor mut sensor mRNAs, miR-19a or miR-Ct mimics and DsRED mRNA. Left, miR-19a presence reduces the embryo specific fluorescence in the presence of wt3?UTR of s1pr1 and only the yolk autofluorescence is visible; right, DsRED expression is constant in all the experiment (G) quantification of the in vivo sensor assay. About 20 embryos were analyzed for each experiment.